Deciphering and engineering microbial metabolic interactions for host and environmental health.

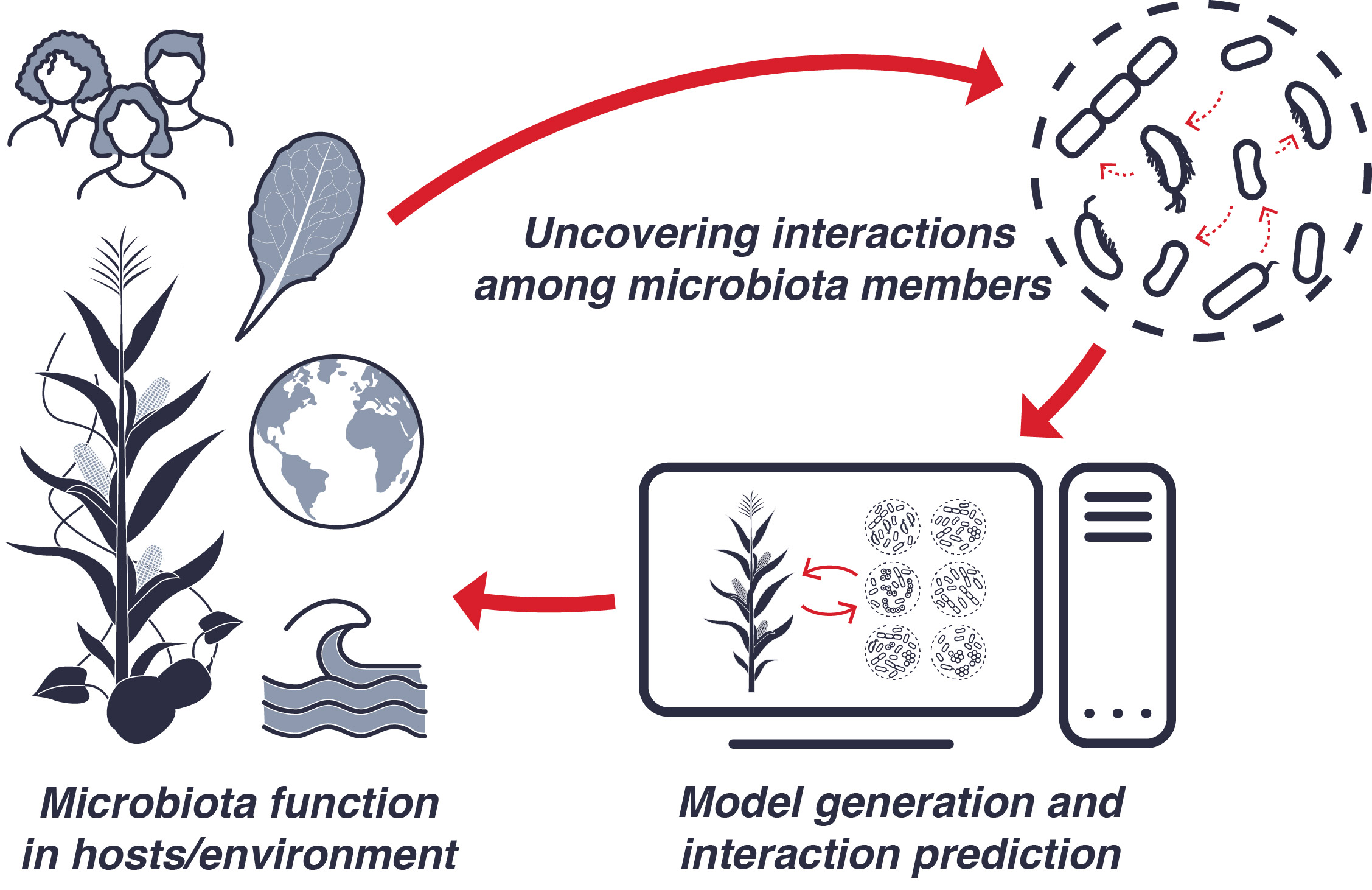 Microbiomes are instrumental to the function of all ecosystems on Earth. An ability to reliably engineer these communities by modulating their structure and function holds tremendous promise to advance applications ranging from bioremediation to enhanced plant, animal, and human health. However, a detailed and general understanding of why certain combinations of microbes are beneficial to their environments while others are not has remained largely out of reach.
Microbiomes are instrumental to the function of all ecosystems on Earth. An ability to reliably engineer these communities by modulating their structure and function holds tremendous promise to advance applications ranging from bioremediation to enhanced plant, animal, and human health. However, a detailed and general understanding of why certain combinations of microbes are beneficial to their environments while others are not has remained largely out of reach.
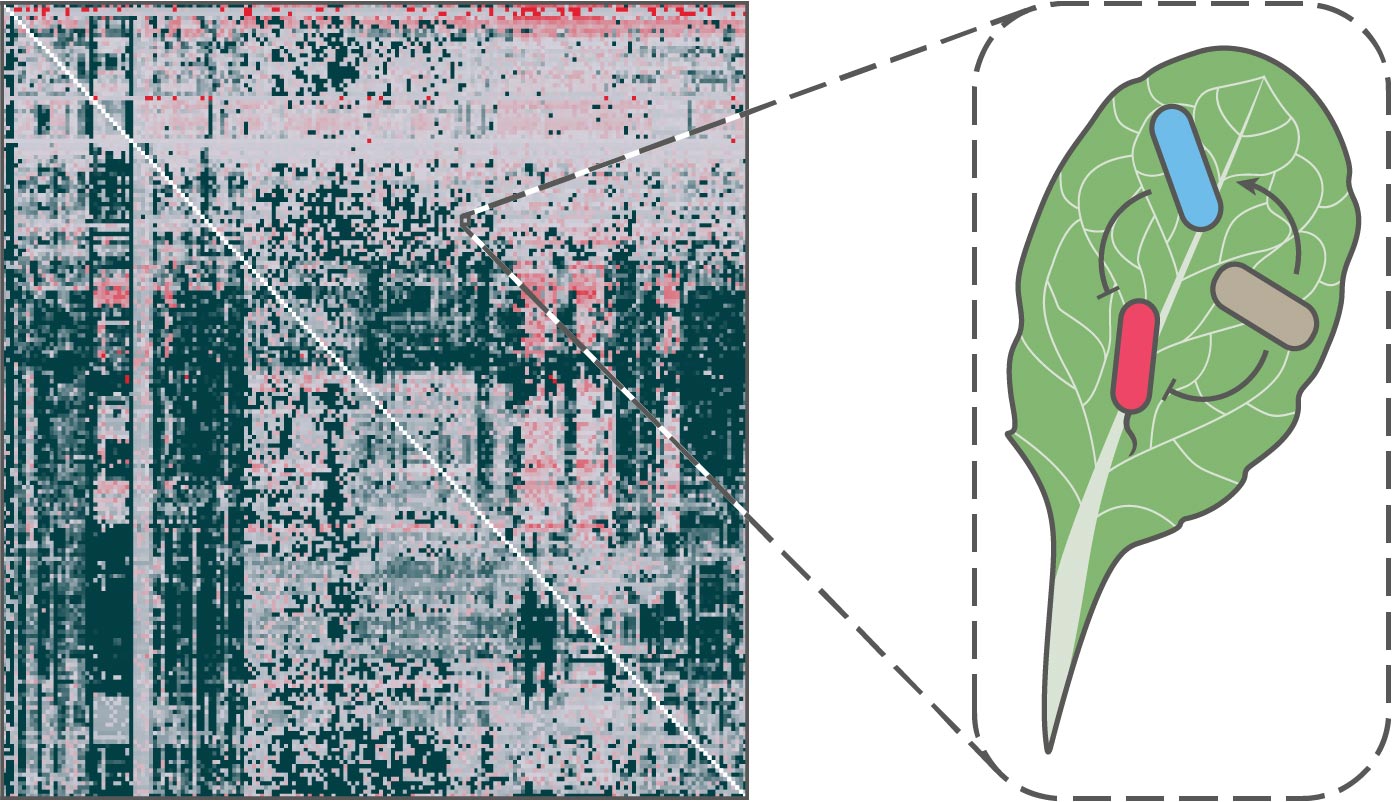
This challenge stems in part from the sheer complexity of microbiomes, whose members engage in a multitude of different interactions with each other and their hosts. However, these interaction networks present a key way to understand the relationship between microbiomes and their host environments. If we can uncover the properties of these interactions and predict their outcomes, we could advance ways to selectively enhance the function of beneficial microbes for improved host and environmental health.
Computational and mathematical models are invaluable tools for understanding interactions in microbiomes. Paired with carefully designed experimental systems, they can shed light on non-intuitive patterns at large scale, contextualize host-microbial evolution, and help reveal the structure and dynamics of complex interaction networks. As such, they are critical for efforts to engineer microbiomes: instead of undertaking a process of trial and error, insights gained from modeling can help efficiently guide the rational design of communities.
The Pacheco Lab pairs computational modeling tools with laboratory experiments and data analysis to understand metabolic interactions in host-associated and environmental microbial communities. Beyond gaining fundamental insights into how metabolic interactions shape community ecology, we seek to apply this knowledge to engineer microbiomes to promote health across hosts. To this end, our research currently focuses on the following complementary themes: